[Study Note] Privacy-Preserving Record Linkage (PPRL) & ML Attacks
What is Privacy-Preserving Record Linkage? (PPRL)
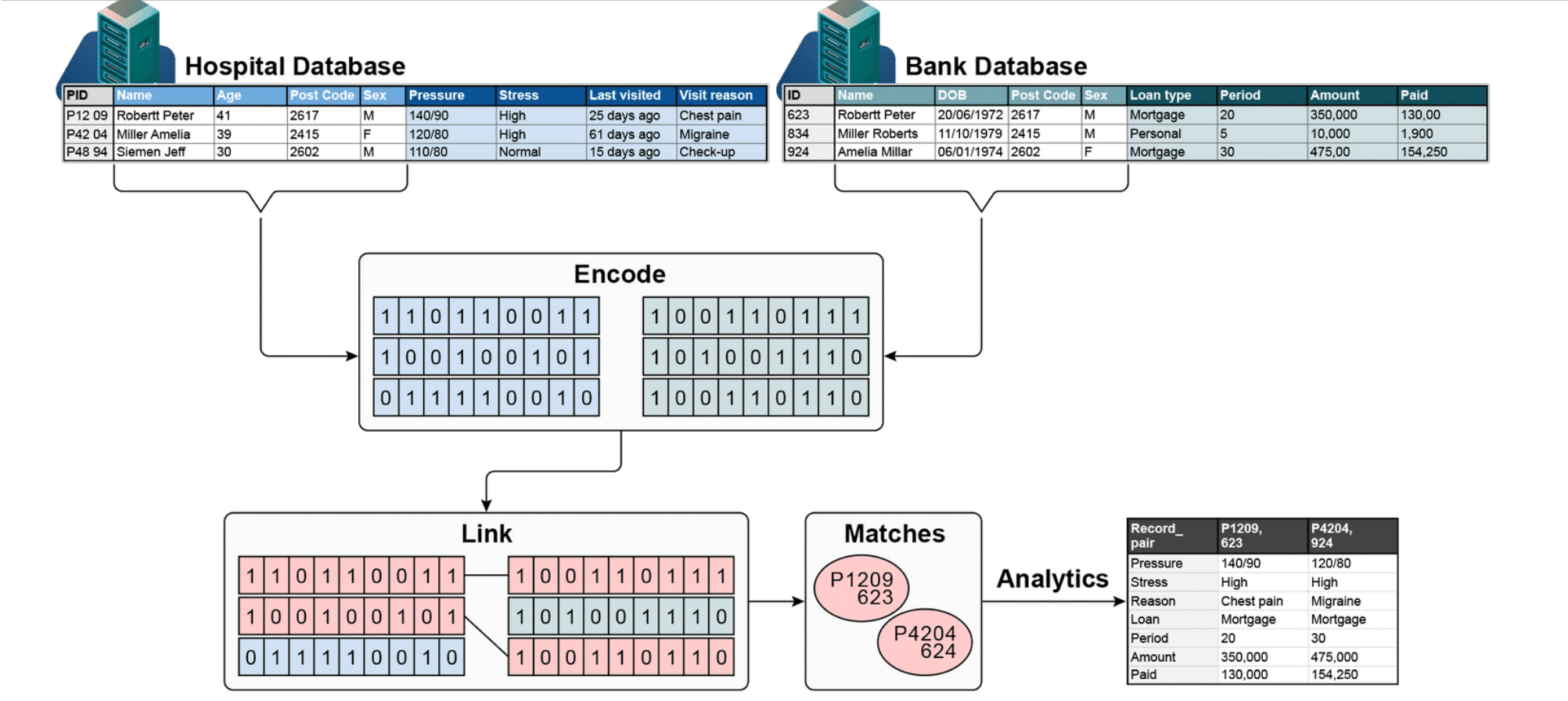
PPRL is a technique for matching records about the same individuals across different databases — without exposing their private data.
How it works
1. Two separate databases exist
- A Hospital DB with patient info (name, age, pressure, etc.)
- A Bank DB with financial info (name, DOB, loan type, etc.)
2. Encode
Instead of sharing raw personal data, each database converts its records into binary vectors (0s and 1s) using a technique like Bloom filters. The actual names/details are never directly shared.
3. Link
The encoded vectors from both databases are compared against each other to find similar patterns.
4. Matches
Records that are sufficiently similar are identified as referring to the same person (e.g., P1209 (hospital) = 623 (bank)).
5. Analytics
Once matched, the combined data can be used for analysis — without either party ever seeing the other’s raw records.
Why it matters
The key privacy guarantee is that neither database sees the other’s sensitive information. Only the encoded representations are exchanged, protecting individual privacy while still enabling useful data integration.
Application of PPRL
A real-world application in the healthcare domain:
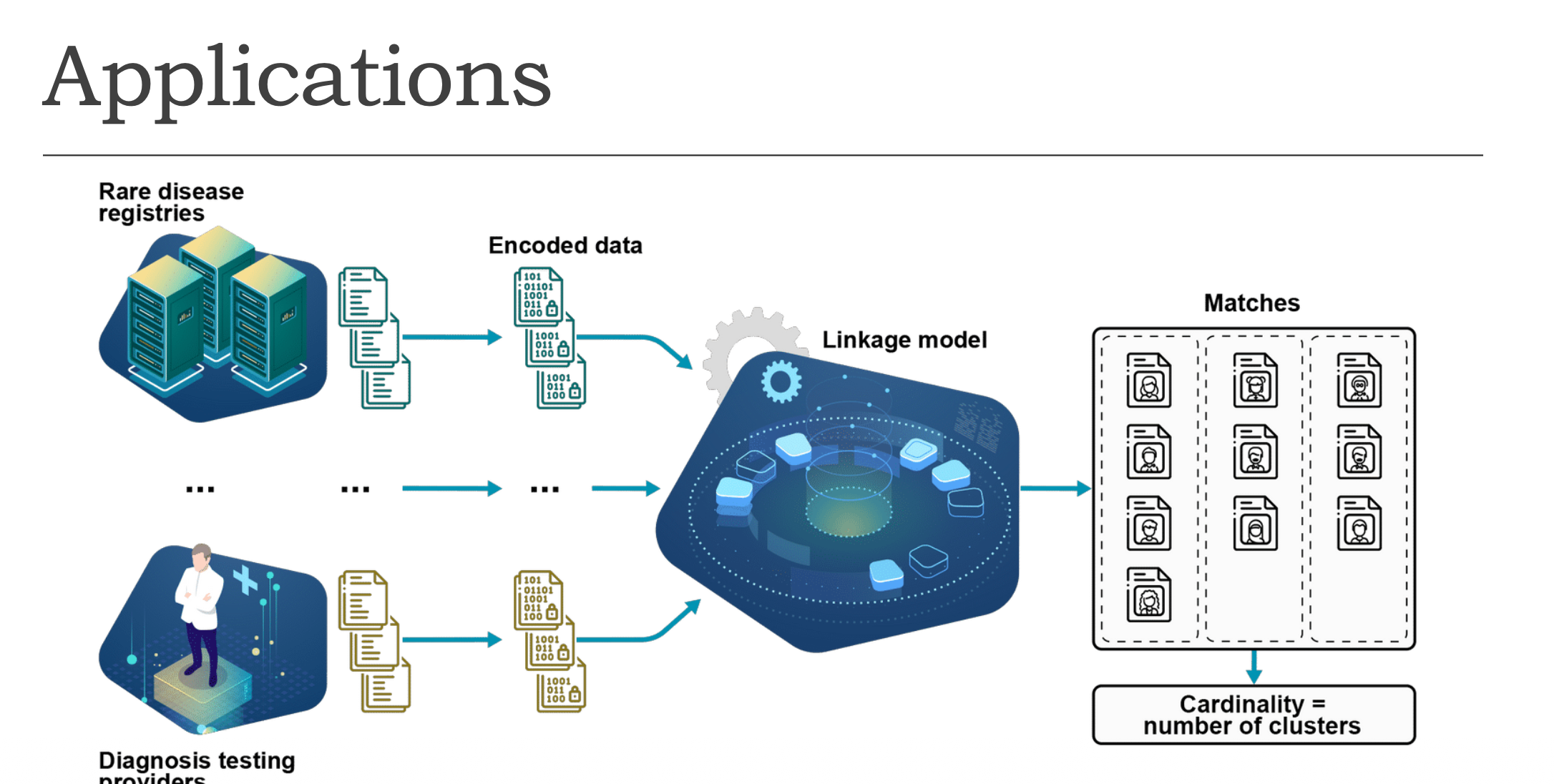
① Two data sources
- Rare disease registries — databases storing records of patients with rare diseases
- Diagnosis testing providers — organisations that conduct medical tests
② Encoding
Both sources convert their raw records into encoded (binary) data before sharing.
③ Linkage Model
All encoded data flows into a central linkage model that compares records across multiple sources.
④ Matches & Clustering
The model groups matched records into clusters — each cluster represents one unique individual. Cardinality = number of clusters (total count of distinct individuals identified).
Key Challenges
- Privacy
- Linkage quality
- Computational efficiency
PPRL Pipeline
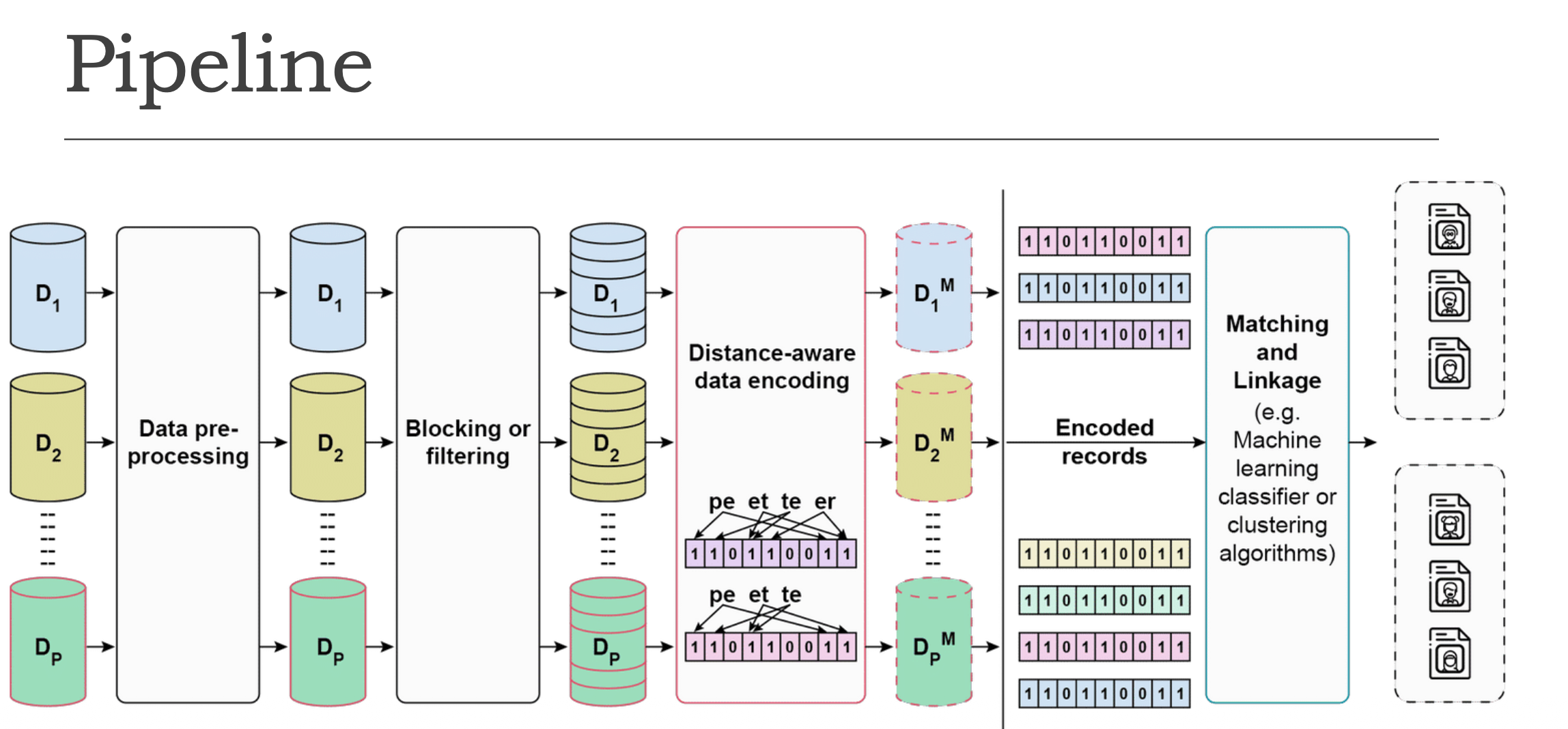
① Multiple Databases (D₁, D₂, … Dₚ) — Several databases from different organisations, each holding raw sensitive personal data.
② Data Pre-processing — Raw data is cleaned and standardised (e.g., fixing typos, formatting names/dates consistently).
③ Blocking or Filtering — Reduces the number of record pairs that need to be compared. Only records that are likely matches are kept.
④ Distance-aware Data Encoding (the key step)
Records are converted into binary vectors (0s and 1s). The encoding preserves similarity — similar records in real life will have similar binary vectors, making them candidates worth comparing more closely.
e.g., “peter” → bigrams (pe, et, te, er) → mapped into a bit vector
⑤ Encoded Records (D₁ᴹ, D₂ᴹ, … Dₚᴹ) — All databases now hold only encoded versions of their data.
⑥ Matching and Linkage — Encoded records are compared using ML classifiers or clustering algorithms.
⑦ Output — Clusters — Final result: groups of matched records, each cluster representing one unique individual.
Blocking
Why do we need Blocking?
When linking two databases, you’d normally compare every record against every other record — this is quadratic complexity. If each database has 1,000 records, you’d need 1,000,000 comparisons. That’s extremely slow and inefficient!
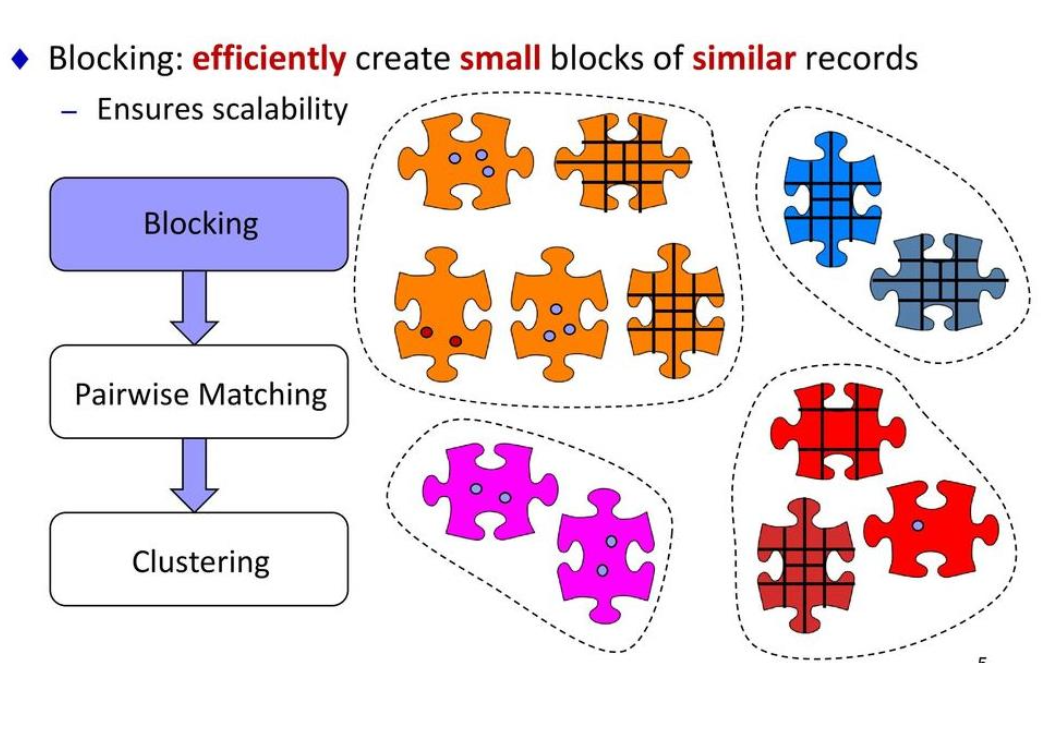
What is Blocking?
Blocking is a filtering step that groups records into smaller buckets called blocks, so you only compare records within the same block.
Example — Postcode as a Blocking Key
| Record | Name | Postcode |
|---|---|---|
| A | John Smith | 2000 |
| B | Jane Doe | 3000 |
| C | Jon Smyth | 2000 |
- Records A and C → same block (postcode 2000) → compared ✅
- Records A and B → different blocks → skipped ❌
Each colour in the puzzle represents a block — only pieces of the same colour get compared.
After Blocking
1
Blocking → Pairwise Matching → Clustering
Blocking dramatically reduces the number of comparisons needed, making the whole linkage process fast and scalable.
Data Encoding: Exact Matching Using Bloom Filters
What is a Bloom Filter?
A Bloom filter is a bit vector (a row of 0s and 1s) that represents a piece of data. Each value gets mapped to specific positions in the vector.
Example: “smith” vs “smith” (identical)
1
2
3
4
5
6
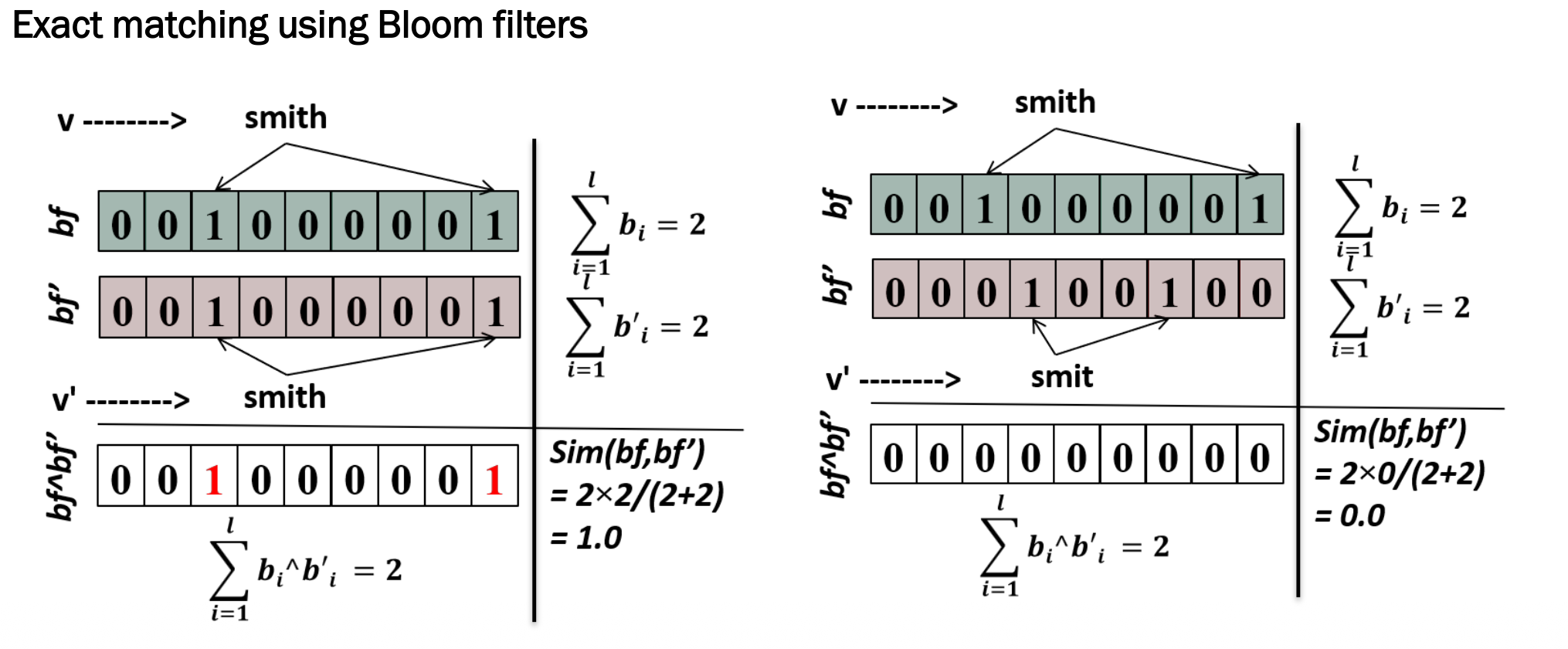
bf = 0 0 1 0 0 0 0 0 1 (smith)
bf' = 0 0 1 0 0 0 0 0 1 (smith)
AND: 0 0 1 0 0 0 0 0 1 → matching bits = 2
Sim = 2 × 2 / (2 + 2) = 1.0 ✅ Perfect match!
Example: “smith” vs “smit” (different)
1
2
3
4
5
6
bf = 0 0 1 0 0 0 0 0 1 (smith)
bf' = 0 0 0 1 0 0 1 0 0 (smit)
AND: 0 0 0 0 0 0 0 0 0 → matching bits = 0
Sim = 2 × 0 / (2 + 2) = 0.0 ❌ No match!
The Similarity Formula (Dice Coefficient)
\[Sim(bf, bf') = \frac{2 \times |bf \wedge bf'|}{|bf| + |bf'|}\]This is called exact matching — even a tiny difference (e.g. “smith” vs “smit”) results in a similarity score of 0.0.
Data Encoding: Fuzzy Matching Using Bloom Filters
String Example: “smith” vs “smit”
Step 1: Break into bigrams
1
2
"smith" → sm, mi, it, th (4 bigrams)
"smit" → sm, mi, it (3 bigrams, missing "th")
Step 2: Map into Bloom filters
1
2
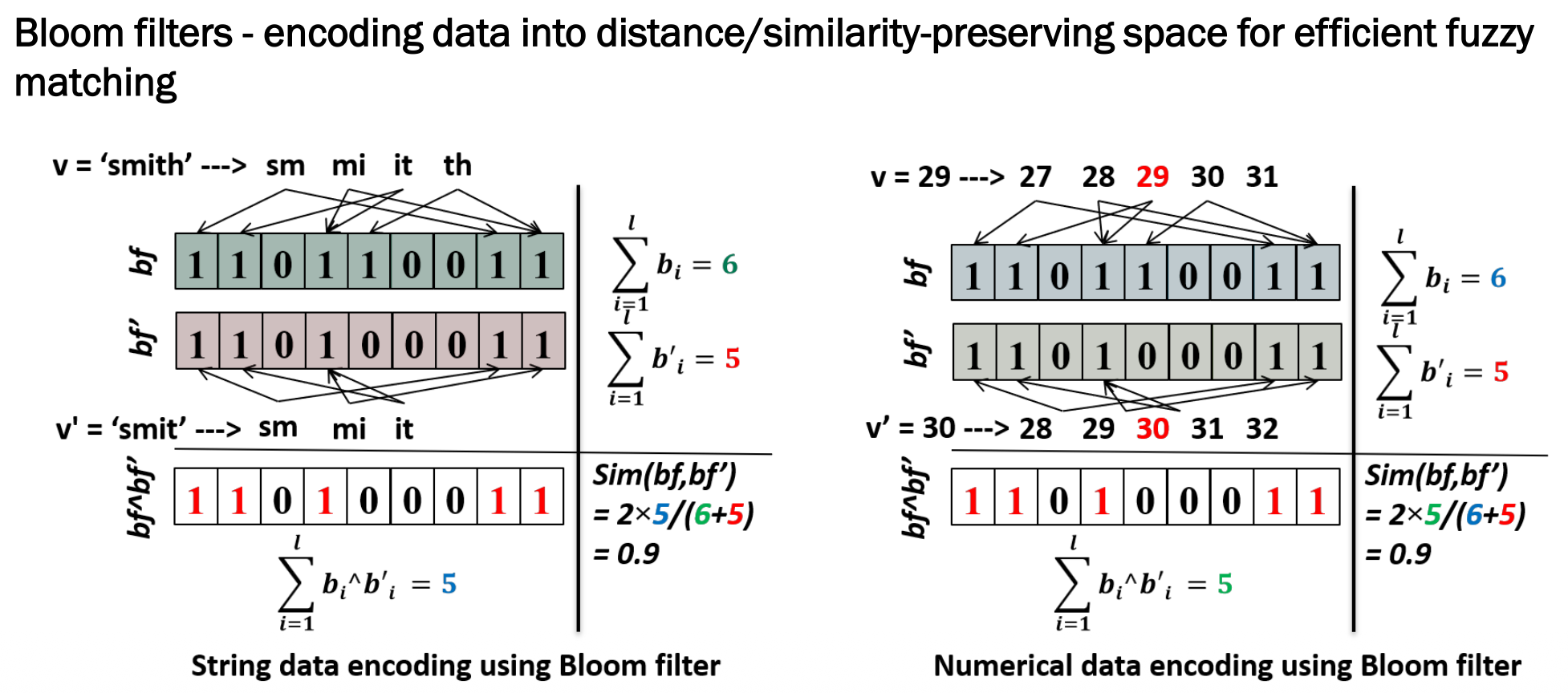
bf = 1 1 0 1 1 0 0 1 1 (6 bits set)
bf' = 1 1 0 1 0 0 0 1 1 (5 bits set)
Step 3: Calculate similarity
1
2
AND = 1 1 0 1 0 0 0 1 1 (5 matching bits)
Sim = 2 × 5 / (6 + 5) = 0.9 ✅ Very similar!
Numerical Example: 29 vs 30
Instead of bigrams, nearby values are generated to capture closeness:
1
2
3
4
v = 29 → {27, 28, 29, 30, 31}
v' = 30 → {28, 29, 30, 31, 32}
Sim = 2 × 5 / (6 + 5) = 0.9 ✅ Very similar!
Key Difference from Exact Matching
| Exact Matching | Fuzzy Matching | |
|---|---|---|
| “smith” vs “smit” | 0.0 ❌ | 0.9 ✅ |
| 29 vs 30 | 0.0 ❌ | 0.9 ✅ |
Fuzzy matching is far more practical in real-world scenarios where typos, abbreviations, and minor errors are common.
Bloom Filters with Provable Privacy Guarantees
The Problem
Bloom filters alone are not fully secure. Attackers can potentially:
- Use cryptanalysis to reverse-engineer original data from the bit vector
- Use pattern mining to identify individuals by analysing repeated patterns
The Solution: Differential Privacy (DP) Noise
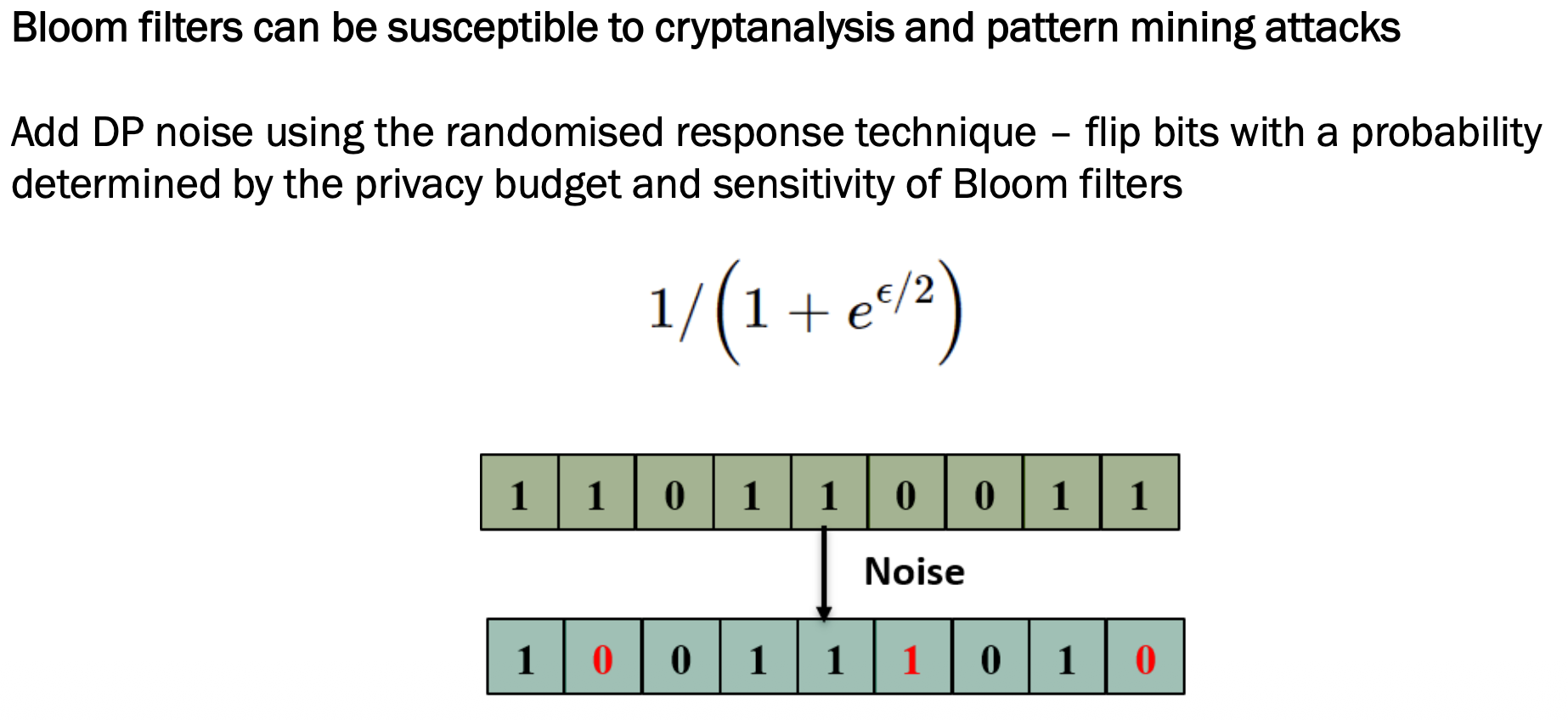
Random noise is added using Randomised Response — randomly flip some bits so attackers cannot fully trust what they see.
Original Bloom filter:
1
1 1 0 1 1 0 0 1 1
After adding noise (flipped bits):
1
1 0 0 1 1 1 0 1 0
The Flipping Probability Formula
\[p = \frac{1}{1 + e^{\varepsilon/2}}\]Where ε (epsilon) is the privacy budget:
| ε value | Flip probability | Privacy | Accuracy |
|---|---|---|---|
| Small ε | High (more flips) | Stronger 🔒 | Lower |
| Large ε | Low (fewer flips) | Weaker | Higher ✅ |
This is the classic privacy vs accuracy trade-off in differential privacy.
Why Attackers Can’t Simply Reverse It
The attacker knows the formula, but NOT which bits were flipped, how many, or where. The flipping is random each time — even running the same name through twice gives different noisy outputs.
Text Data Encoding: Counting Bloom Filters (CBF)
Why a Different Approach for Text?
Regular Bloom filters just store 0s and 1s, which works for names and numbers. But for longer medical text, we need to capture how important each word is — not just whether it appears. That’s where Counting Bloom Filters (CBF) come in.
Step 1: TF-IDF Weighting
| Record v | Score | Record v’ | Score |
|---|---|---|---|
| biopsy | 3 | cancer | 5 |
| cancer | 6 | polyp | 3 |
| mass | 1 | biopsy | 4 |
| polyp | 1 | tumour | 2 |
Step 2: Map into CBF
1
2
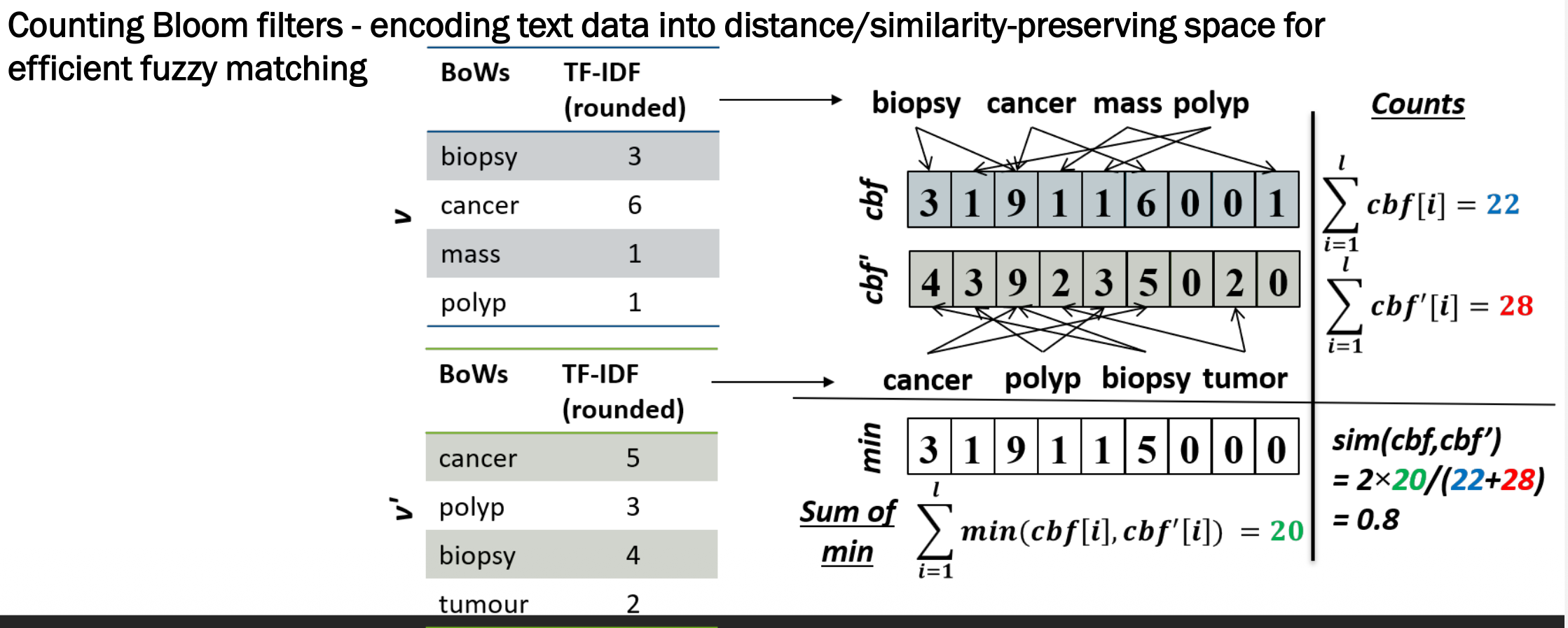
cbf = 3 1 9 1 1 6 0 0 1 (total = 22)
cbf' = 4 3 9 2 3 5 0 2 0 (total = 28)
Step 3: Calculate Similarity
1
2
min = 3 1 9 1 1 5 0 0 0 (sum of min = 20)
sim = 2 × 20 / (22 + 28) = 0.8 ✅ Pretty similar!
A score of 0.8 correctly reflects that the two records share cancer, biopsy, and polyp but differ in “mass” vs “tumour” — very similar but not identical.
Matching and Linkage
| Method | Pros | Cons |
|---|---|---|
| Threshold-based | Simple, fast | Hard to set the right threshold |
| Probabilistic (Fellegi-Sunter) | Accounts for errors | Needs prior error estimates |
| Rule-based (Deterministic) | Interpretable | Time consuming to build |
| ML-based | Flexible, accurate | Needs labelled training data |
Evaluation
1. Computational Efficiency
| Metric | What it measures |
|---|---|
| Runtime | How long it takes to run |
| Memory consumption | How much RAM is used |
| Communication size | How much data is sent between databases |
| Reduction ratio | How much blocking reduced comparisons |
| Pairs completeness | Whether blocking kept all true matches in same block |
2. Linkage Quality
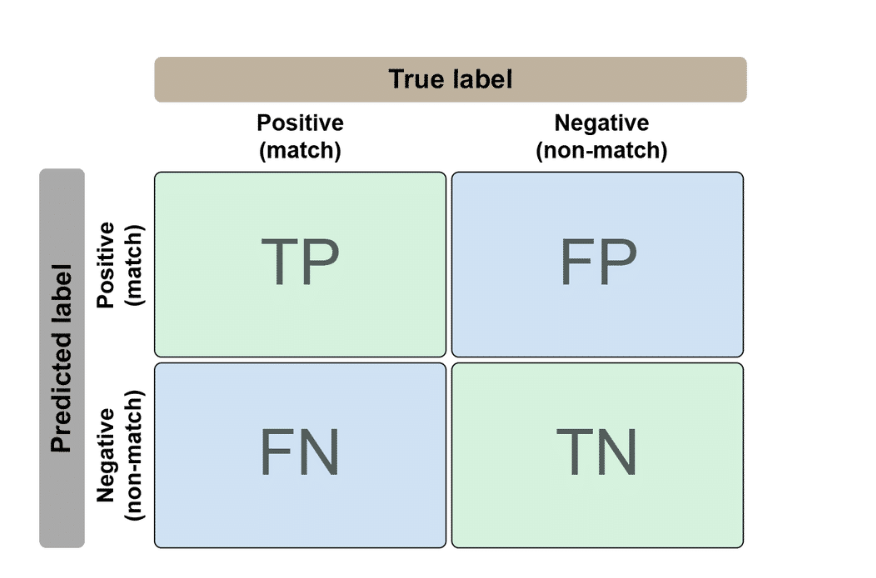
| Actually a Match | Actually NOT a Match | |
|---|---|---|
| Predicted Match | TP (correct!) | FP (false alarm) |
| Predicted Non-match | FN (missed!) | TN (correct!) |
3. Privacy
| Metric | What it measures |
|---|---|
| Entropy / information gain | How much info an attacker could extract |
| False positive rate of Bloom filter | How hard it is to reverse-engineer original data |
| Privacy budget (ε) | How much noise was added via differential privacy |
Machine Learning: Attacks on ML Models
AI and ML models can be attacked in two main ways:
1. Inference Attacks
Membership Inference Attack — “Was this specific person’s data used to train this model?”
Attribute Inference Attack — “What are this person’s private attributes?” (uses the model to fill in missing info)
2. Adversarial Attacks
The attacker manipulates the input to trick the model into producing a desired (incorrect) output.
| Attack type | Goal |
|---|---|
| Membership inference | Find out if a record was in the training data |
| Attribute inference | Infer private details about an individual |
| Adversarial | Manipulate the model’s output |
Membership Inference Attack (MIA)
Given a trained ML model and a record, determine whether that record was part of the model’s training data.
How it works
1
2
3
4
5
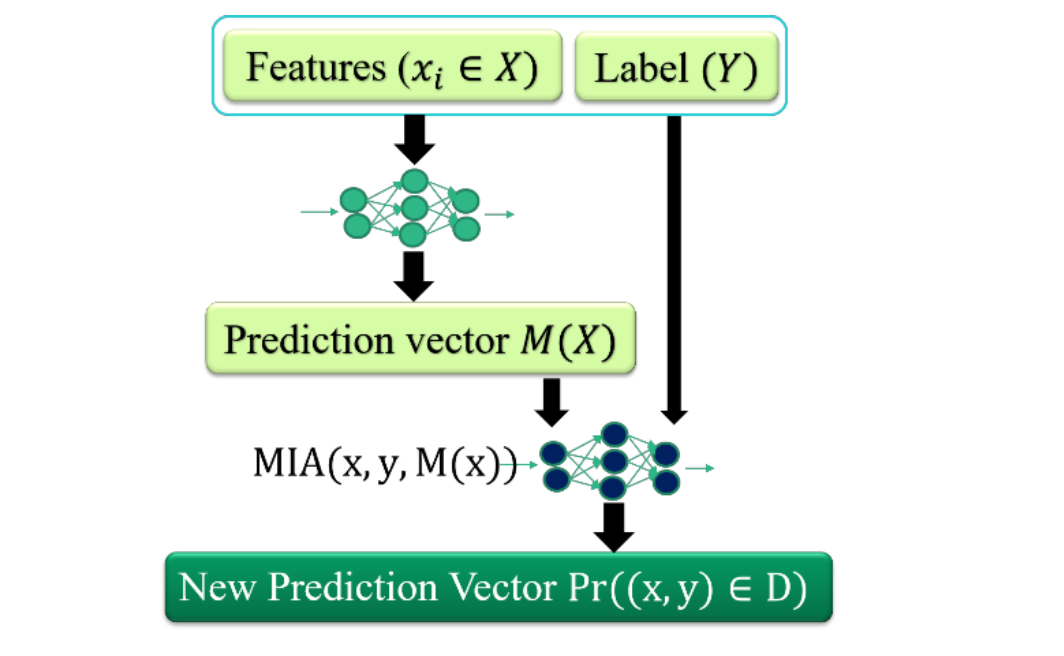
Features (x) + Label (y) → ML model → Prediction vector M(x)
↓
MIA(x, y, M(x))
↓
Pr((x,y) ∈ D) → Member or Not?
Why does this work?
| Data type | Model behaviour |
|---|---|
| Training data (member) | Very high confidence, low loss |
| New data (non-member) | Lower confidence, higher loss |
The attacker exploits this difference to distinguish members from non-members.
Black Box MIA
In a black box setting, the attacker can only send inputs and observe outputs — no access to model weights, architecture, or training data.
Three Levels of Information
| Setting | Info available | Attack difficulty |
|---|---|---|
| Full confidence scores | All class scores | Easy |
| Top-k confidence scores | Partial scores | Moderate |
| Prediction label only | Just the label | Hard |
Why We Can’t Always Use Label Only
Even with just a label, MIA is still possible (harder, but not impossible). More importantly, in PPRL you need to know how similar two records are — a binary “match / non-match” loses critical information.
Real Defences
- Differential privacy during training
- Output perturbation (adding noise to confidence scores)
- Restricting output detail (label only where possible)
- Limiting query rates
MIA against Text Linkage Model
| Detail | |
|---|---|
| Input | Free-text mentor feedback (sequence of words) |
| Output | Categorical label (feedback classification) |
Standard MIA definitions may not work well here — free-text input requires a looser/relaxed definition of MIA.
Three Black-box MIA Approaches
| Attack | Method | Complexity |
|---|---|---|
| Shadow model (Shokri et al., 2017) | Trains multiple shadow models to mimic target | High |
| Prediction loss (Yeom et al., 2018) | Low loss → likely member | Moderate |
| Max confidence (Salem et al., 2018) | High confidence → likely member | Low |
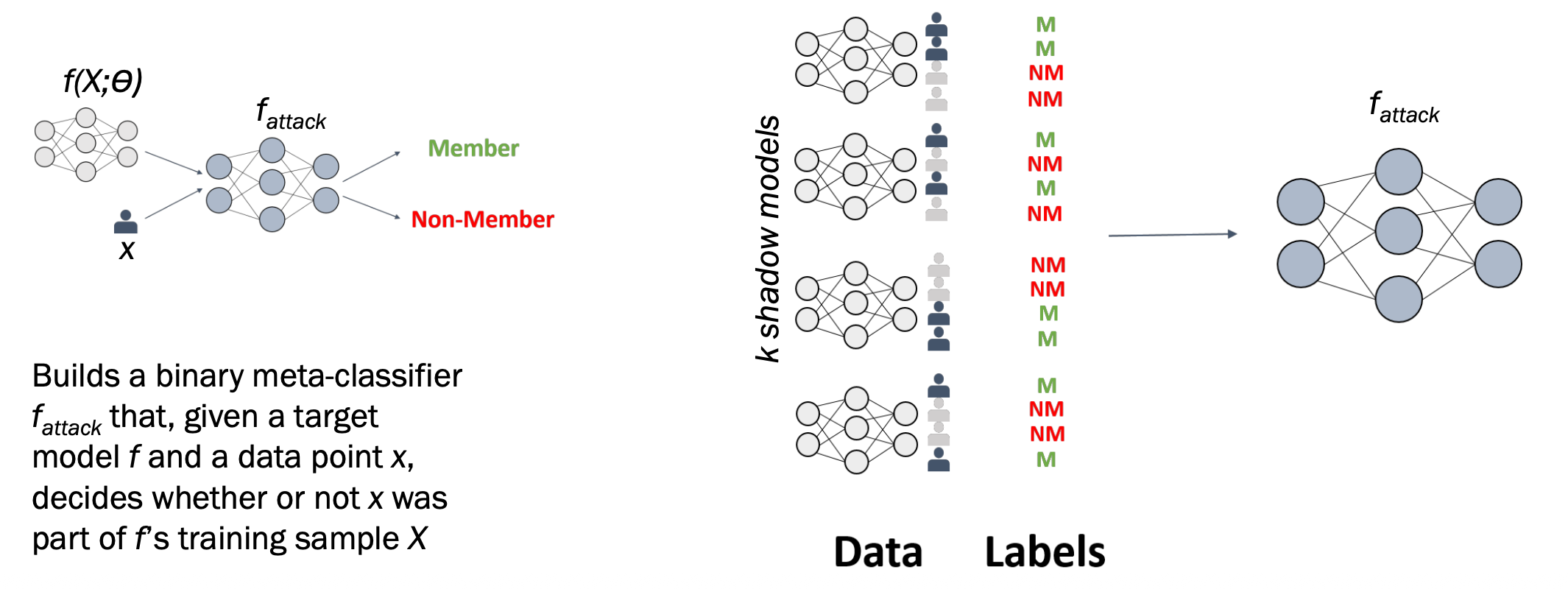
Shadow Model-based MIA
The attacker creates fake versions (shadow models) of the target model to learn how it behaves.
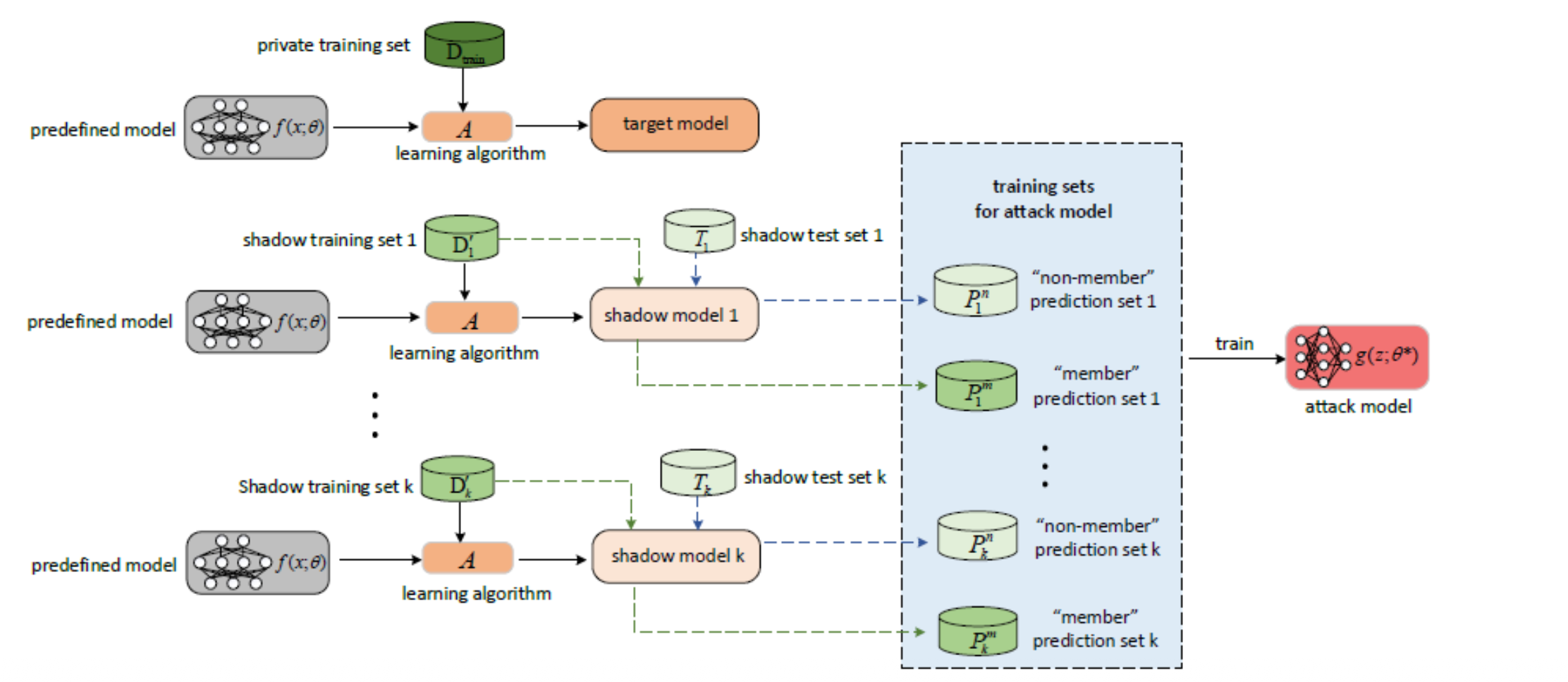
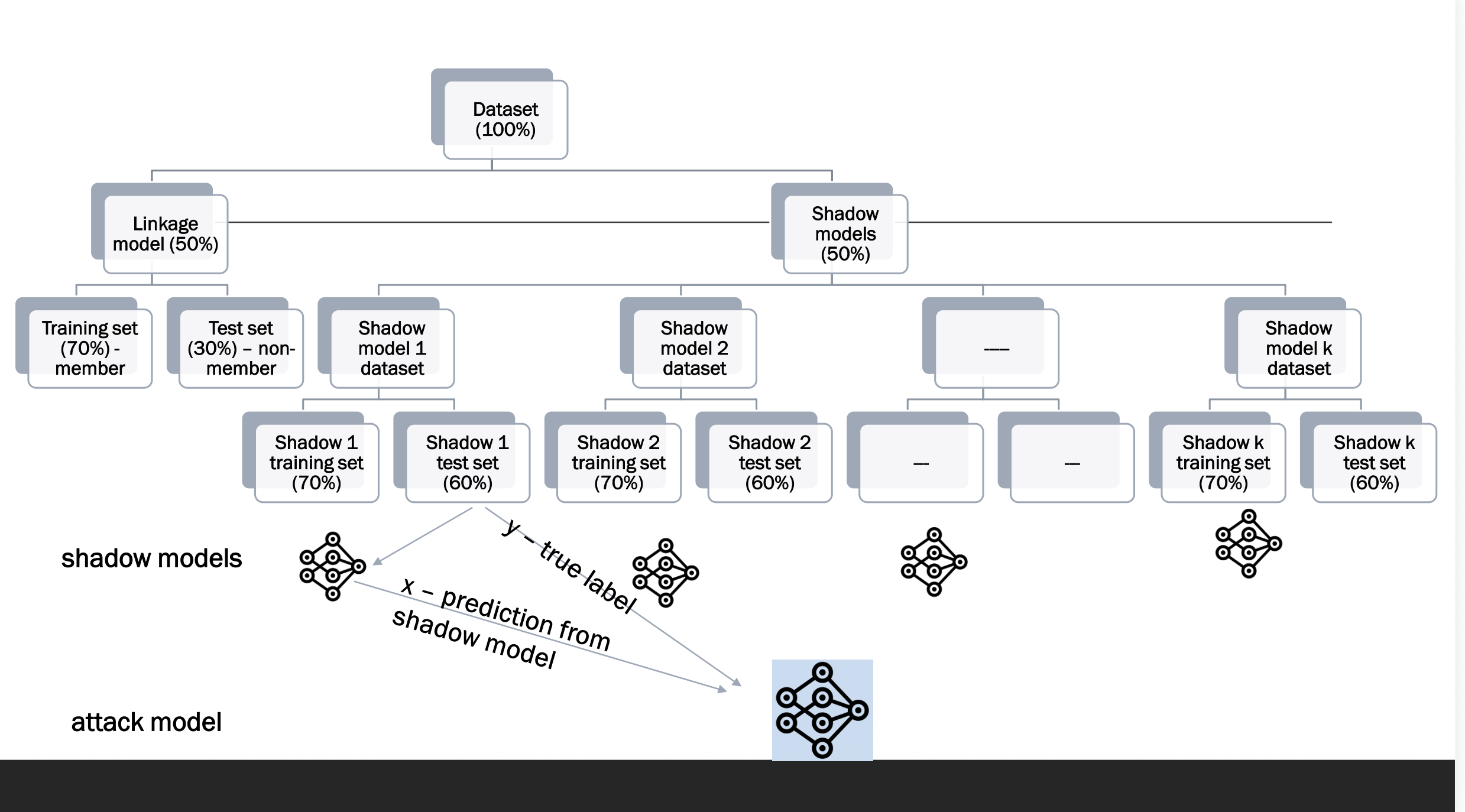
Step-by-Step
Step 1: Target model trained on private data D_min — the attacker cannot see inside it.
Step 2: Attacker trains k shadow models on similar but different data to mimic the target.
Step 3: For each shadow model, the attacker labels outputs as member (M) or non-member (NM).
Step 4: All labelled outputs train the attack model f_attack.
1
2
3
4
5
6
7
8
9
Shadow models learn to mimic target model
↓
Generate M / NM labelled predictions
↓
Train f_attack on those labels
↓
Apply f_attack to the real target model
↓
Determine: Member or Non-Member? 🔍
Analogy: Like practising on past exam papers to learn the examiner’s patterns, then applying that knowledge to the real exam.
Experiments: MIA with Standard Input Space (CIFAR-10)
| Detail | Value |
|---|---|
| Dataset | CIFAR-10 image dataset |
| Data size | 6,000 images, 10 classes |
| Image size | 32×32 pixels (RGB) |
| Model | Logistic regression |
| Attacks | Threshold-based + Shadow model-based |
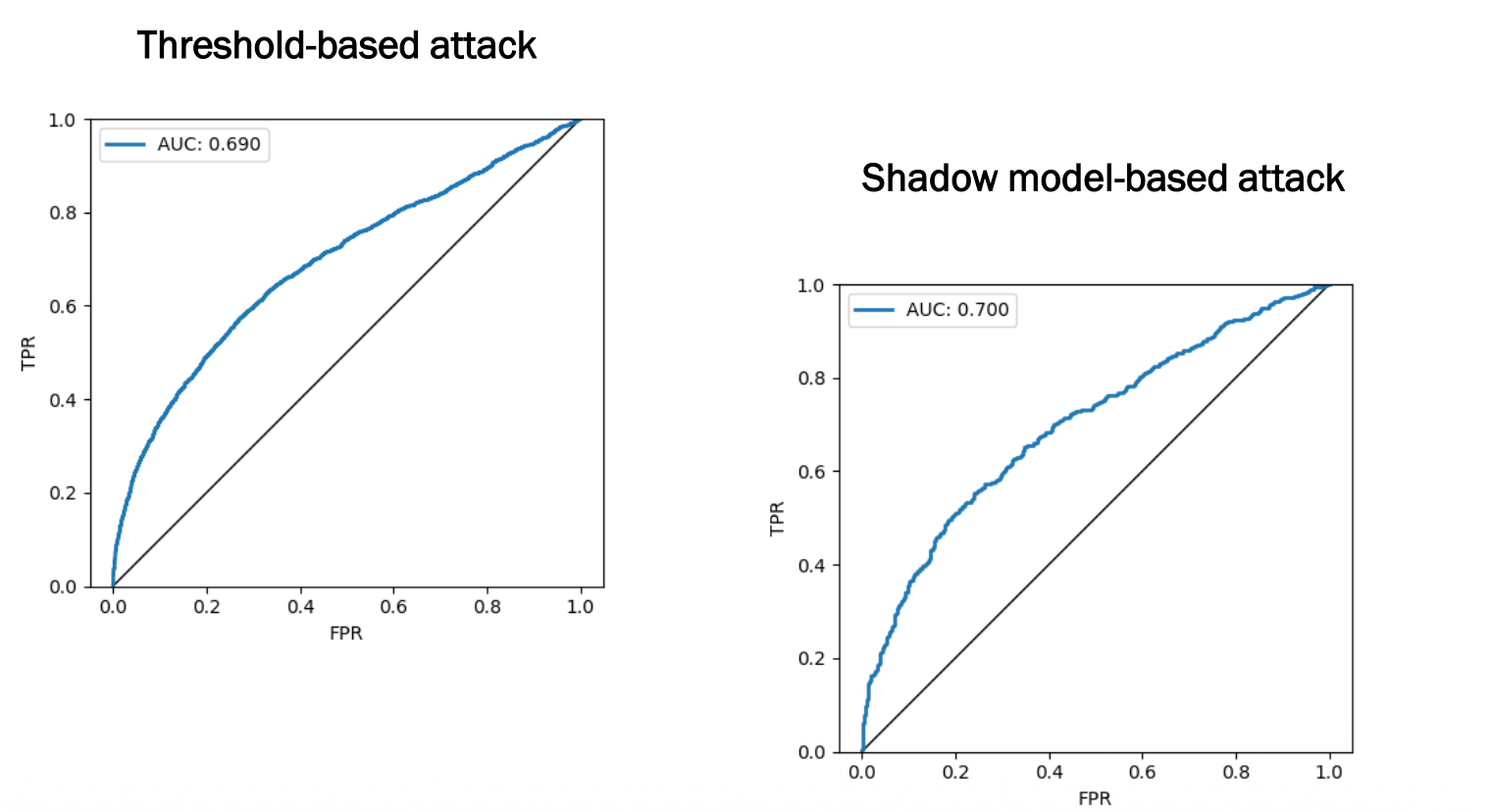
| Attack | AUC |
|---|---|
| Threshold-based | 0.690 |
| Shadow model-based | 0.700 |
Both attacks are meaningfully better than random guessing (AUC = 0.5), confirming that even a simple logistic regression on image data leaks membership information.
Experiments: MIA with Free-text Input Space
| Detail | Value |
|---|---|
| Dataset | Free-text mentor feedback |
| Pre-processing | Remove stop words + lemmatise |
| Representation | Bag of Words (2,233 BoWs) |
| Labels | 5 categories (outstanding → poor) |
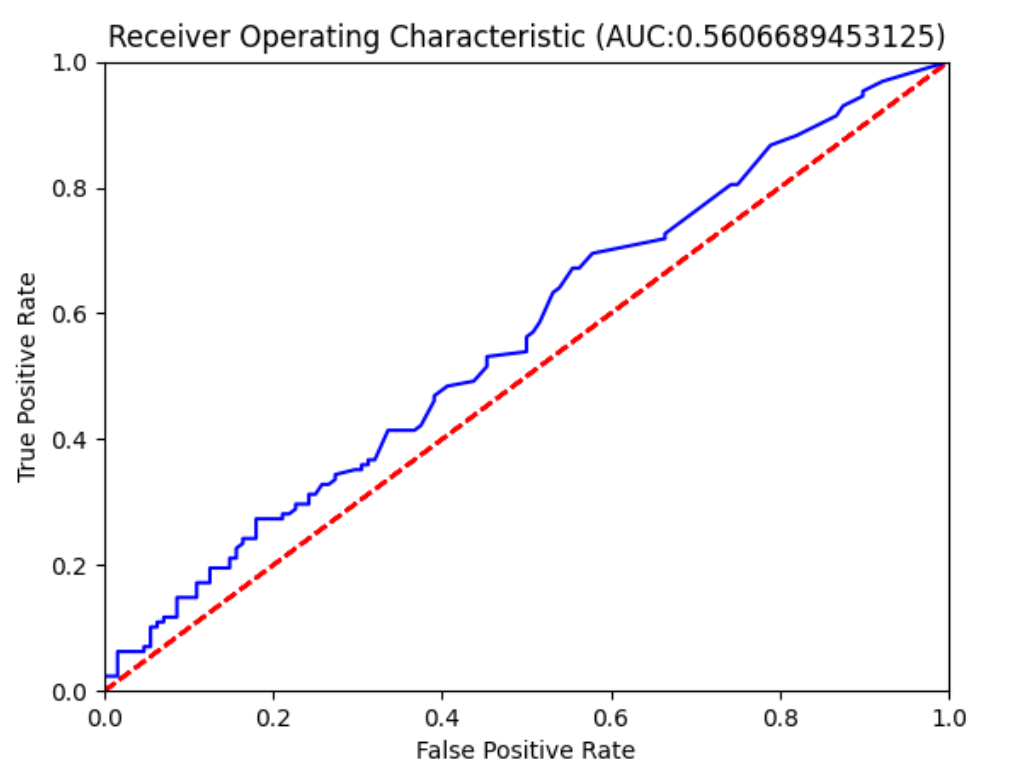
| Model | Logistic regression |
Result: AUC = 0.56 — barely better than random guessing.
Why is the AUC So Much Lower Than CIFAR-10?
- High dimensionality — 2,233 sparse BoW features vs clean RGB pixels
- More variability — people write feedback many different ways → less overfitting
- 5 output classes — confidence scores more spread out, harder to exploit
| Setting | Attack | AUC |
|---|---|---|
| CIFAR-10 | Threshold | 0.690 |
| CIFAR-10 | Shadow model | 0.700 |
| Free-text (large) | Shadow model | 0.560 |
Experiments: Free-text MIA (Small Dataset, Overfitting)
| Detail | Value |
|---|---|
| Dataset | Small free-text feedback (1,633 BoWs) |
| Model | ANN classifier |
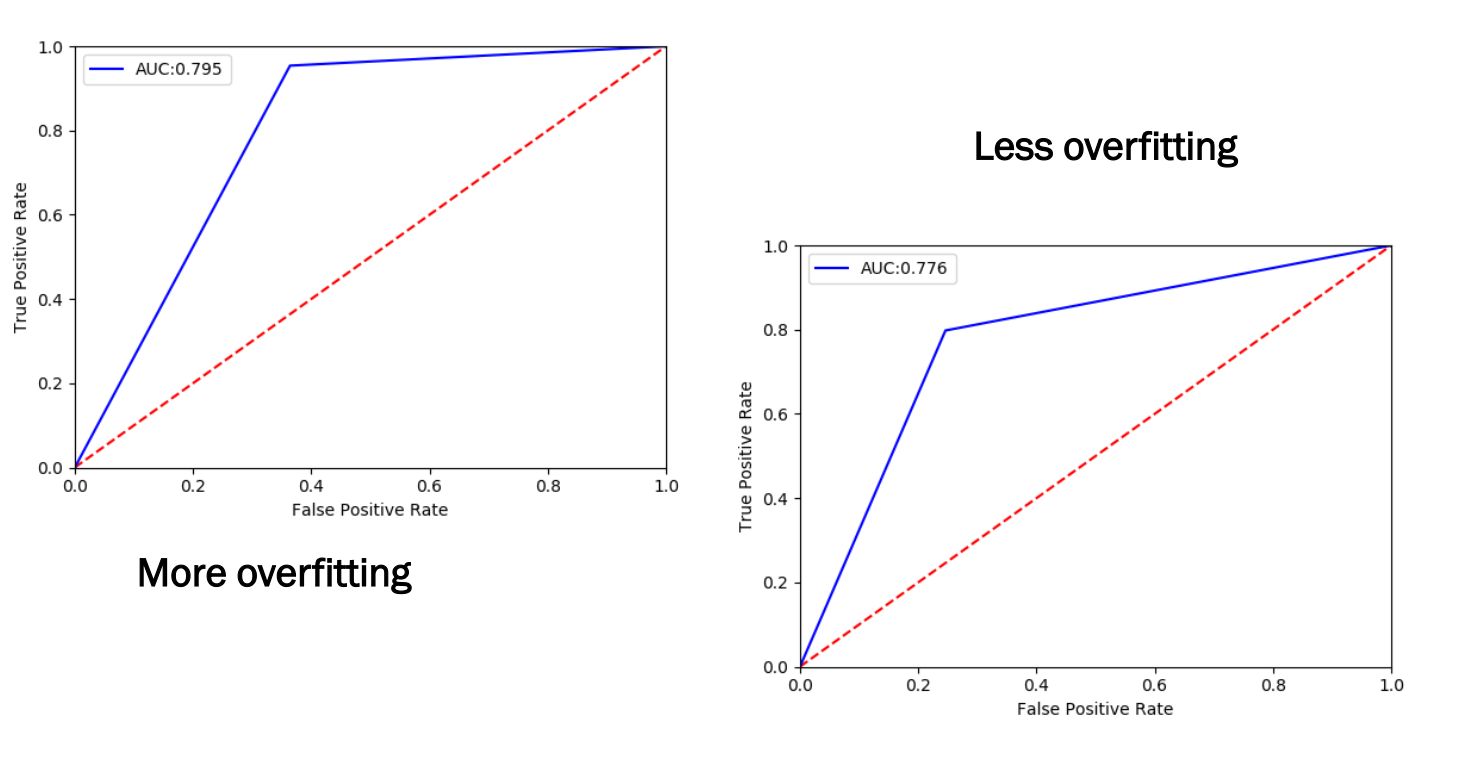
| Setting | AUC |
|---|---|
| More overfitting | 0.795 |
| Less overfitting | 0.776 |
Why Does Overfitting Make MIA Easier?
1
2
3
4
5
6
Overfitted model:
Members → Very high confidence, very low loss
Non-members → Much lower confidence, higher loss
More overfitting → Bigger gap → AUC 0.795 (easier to attack)
Less overfitting → Smaller gap → AUC 0.776 (slightly harder)
Reducing overfitting (regularisation, dropout, early stopping) is an effective privacy defence as well as good ML practice!
Experiments: Free-text MIA (Multiple Shadow Models)
| Number of shadow models | AUC |
|---|---|
| 10 shadow models | 0.776 |
| 5 shadow models | 0.747 |
More shadow models → more diverse training data for the attack model → higher AUC.
Full Summary of Free-text Experiments
| Setting | AUC |
|---|---|
| Large dataset, logistic regression | 0.560 |
| Small dataset, less overfitting, 10 shadow models | 0.776 |
| Small dataset, more overfitting, 10 shadow models | 0.795 |
| Small dataset, less overfitting, 5 shadow models | 0.747 |
Three Key Factors That Increase MIA Vulnerability
- Smaller dataset → more overfitting → easier to attack
- More overfitting → bigger gap between member/non-member behaviour → easier to attack
- More shadow models → better trained attack model → easier to attack